Comma separated Value files also known as CSV files can be imported and exported to and from R. This article explains how to import and export CSV files in R.
Import CSV file in R
First, we are going to discuss the importing of CSV file in R using read.csv and read_csv functions. The most common method for importing a CSV file into R is by using the base R. The base R uses read.csv function to import CSV file.
There are multiple files we are going to use for the practice, thus you can download all these files from below
Download Example FileLet’s say we have a file saved by the name of example_data, and we want to import it into R using the base function R, we use the following command
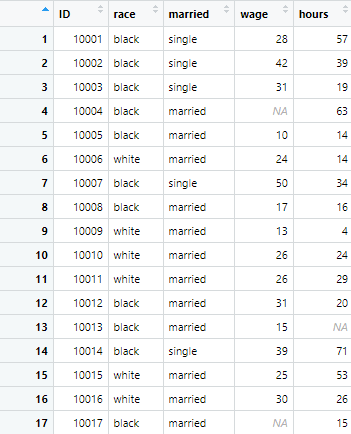
r.c <- read.csv("example_data.csv")The dataset imported into R contains 500 observations across 5 variables

If you look into above data set, there are headers for each column, named ID, race, married etc. Thus, importing such a file is straightforward. However, if we come across a data set that doesn’t have headers, and we use the same command as we did earlier, it will assign first row of observation as the header names to data. Let’s import a file that doesn’t have headers using the following command,
r.c_h1 <- read.csv("noheader.csv")The above command uses observations of first row as headers. This is shown in image below
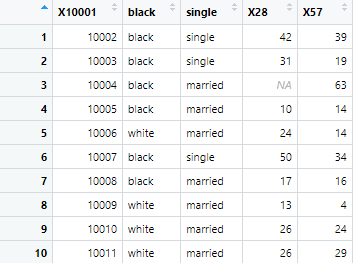
However, the first row of observations don’t represent the column names, and cannot be used as headers. Thus, we need to instruct R, not to use first row of observations for headers. To prevent R from selecting the first row of observations as headers, we use the command given below
r.c_h2 <- read.csv("noheader.csv", header=FALSE)The parameter “header=FALSE” will instruct R not to assign any headers to the data set, and the data set will look like following
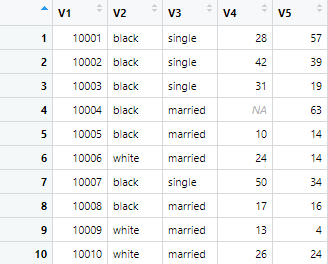
Now, most of the time, we have a comma separated files, having values in each column separately. But there are instances, where instead of comma separator, we have data in a file which is separated by other delimiters such as semicolons. Let’s say we have a file in which data is separated by semicolons. To import such kind of file in to R, we can use the read.csv function in the following way.
r.c_sep1 <- read.csv("colon_seperated.csv")The practice data set can be downloaded from above files saved by the name of colon_seperated. The data of the file looks like following, which makes it hard to further work with data
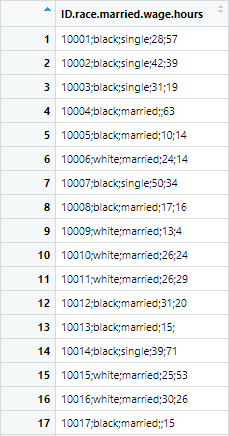
To separate these observations in separate columns, we use the “sep” parameter in R, which will allow R to separate the data currently joined by semicolons. The command for this purpose will be following
r.c_sep2 <- read.csv("colon_seperated.csv", sep = ";")These commands separate the data into distinct columns, replacing missing observations with NA values.
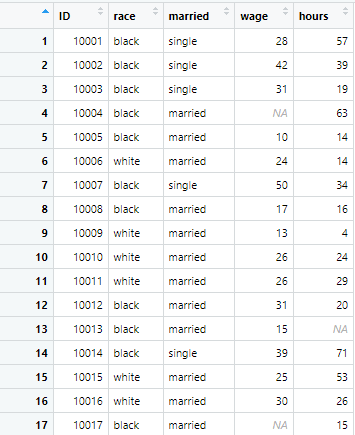
Just like data can be separated from semicolons, it can also be separated from decimals in an Excel file. It is important to note that Excel file can use different kind of decimal separators depending on the settings. In some files, a period or full stop is used as a decimal separator, while others have comma as a separator in them.
For instance, we have a file that has comma separator in its data. To import that file, we use the following command
r.c_dec1 <- read.csv("decimal.csv")The data file is visualized as below. Note that hours variable has data separated by commas in the given data.
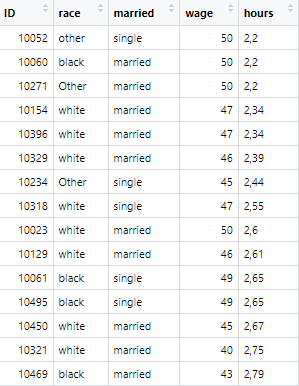
We can format Excel files in such a way that we can add row and column numbers. Or we can also use a column already present in the data set as row names. For instance, in our original data set, the observations of first column, named ID, can be used as row numbers.
To assign the first column as row numbers, we use the following command in R
r.c_name1 <- read.csv("example_data.csv", row.names = 1)The parameter row.names= 1 in above command instructs R to use observations of first column as row names.
Importing CSV file using tidyverse
Till now, we have been working with base R to import Excel files and manipulate these files. However, we can move towards much easier ways to import the Excel files by using the “read_csv” function in the tidyverse package. This is a quicker and more efficient method for importing data in R, specially if the data sets are large.
First, install and load the readr package by using the following commands
install.packages(readr) library(readr)
Next we load the example data set again, but now with a slightly different command, as shown below
r_c <- read_csv("example_data.csv")The data set is same as loaded earlier.
Similarly, as we did earlier , if we have a file without headers, we can use the following command, which will not select the headers by default from observations of first rows.
r_c_h <- read_csv("noheader.csv", col_names = FALSE)Or if we want to separate data joined by semicolons, we use the following commands, which would remove the semicolons from observations in data.
r_c_sep1 <- read_csv("colon_seperated.csv") r_c_sep2 <- read_delim("colon_seperated.csv", delim = ";")The read_csv function is faster and can load the data set containing millions of observations, without crashing or lagging the system. Thus, if you are working with a large data set, it is advised to use read_csv function for importing CSV file in R.
Exporting CSV file from R
CSV file can be also be exported from R. It can be done using the write.csv function, This function is part of R base.
We use the write.csv function in the following way
write.csv(single,"single_data.csv")
The above command will save the filtered data of single people in CSV file in the current working directory.
The data file that we just saved, can also be saved at the location of our choice, instead of in current working directory. For example, if you want to save it to a different folder, you can use the specified file path and the CSV file will be saved in that folder. The example command for this purpose will be following
write.csv(single, "D:\single_data.csv")
Similarly, there are other changes that could be made in file before exporting it. For example, if you want to exclude the name of columns and rows in the file, it can also be done using the write.csv function. To instruct R not to include row and column names in R, we use the following command
write.table(single,"single_data.csv", sep = ",", row.names = FALSE, col.names = FALSE)
By setting the parameters row.names and col.names equal to FALSE, we are instructing R not to include headers and row names in file. The CSV file saved has data that looks like following
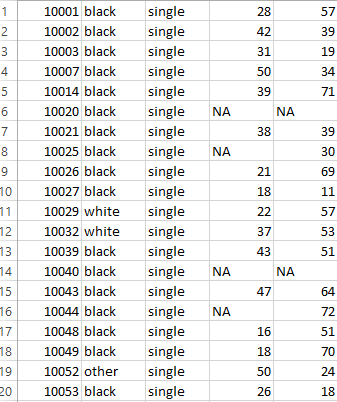
Note that in above image, there are observations represented by “NA” , which means these observations are missing. If we want “NA” to be replaced by the word “missing” or just the blank space or any other character that would represent missing values in our data, we use the following commands
write.csv(single,"single_data.csv", na = "Missing") write.csv(single,"single_data.csv", na = "")
Till now, we discussed exporting the CSV file in R using the base R. However, there is a quicker and easy way to do so by using the package present in tidyverse’s set of packages.
To use that function, we first load the package by using the following command
library(readr)
Like previous operations, if we want to instruct R not to include column names in the file while exporting it, we use the write_csv function in the following way
write_csv(single,"D:\single_data.csv", col_names = FALSE, append = TRUE)
So the above command writes the data without column names and appends it to an existing CSV file (or creates a new one if it doesn’t exist)
Or if you want to replace “NA” values with “missing values” use the write_csv function in the following way
write_csv(single,"D:\single_data.csv", na = "Missing")
The exported CSV file has data in following way
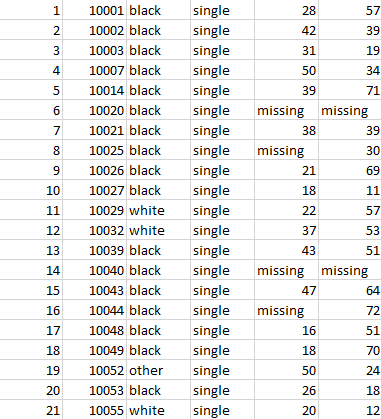
In conclusion, this article has explored the essential techniques for importing and exporting CSV files in R, offering both base R and tidyverse approaches to suit varying data processing needs.
